
Next-Generation Sequencing (NGS) involves high-throughput sequencing technologies that have, over time, become affordable to most labs. The ability to generate large volumes of data has greatly empowered genetic, biological and medical research.
Sequencing technologies involve three main processes in the following order: library preparation, sequencing and analysis. The choice of technology or combination of technologies depends on the purpose of the project; for example, resequencing vs de novo sequencing and assembly will require different approaches.
AgriGenome offers a comprehensive range of next-generation sequencing services utilizing multiple sequencing platforms including Illumina HiSeq, MiSeq, PacBio Sequel and Oxford Nanopore Technology. Our team of scientists and Bioinformaticians work with you to deliver reliable high-quality results and provide end-to-end solutions for your NGS project needs.
Exome sequencing (also known as targeted exome capture) is a strategy to selectively sequence the coding regions of the genome and is a more affordable alternative to whole genome sequencing. In the human genome, there are about 180,000 exons: they make up about 1% of the human genome, which translates to about 30 megabases (Mb) in length.
It is estimated that the protein coding regions of the human genome account for about 85% of the disease-causing mutations. Exome sequencing is a productive and cost effective method to identify functional variations that are responsible for both Mendelian and common diseases.
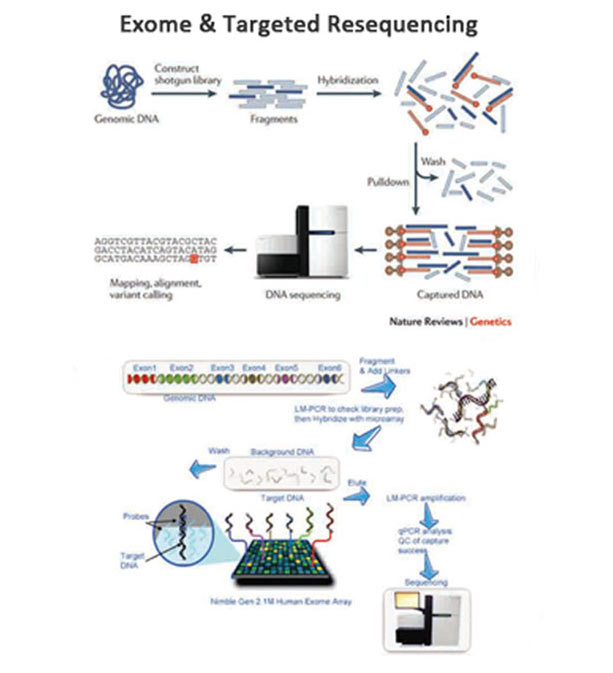
Platform : We offer exome sequencing on Illumina HiSeq.
For sales inquiries, please email us at: sales@aggenome.com
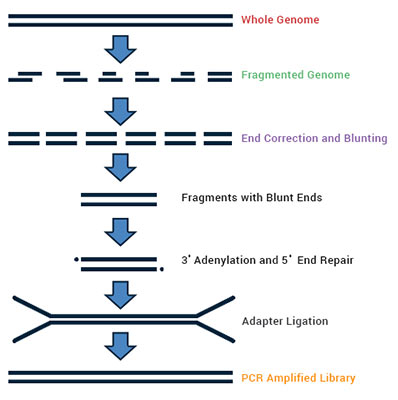
De novo genome sequencing is used to characterize genomes where, either there are no reference sequences available, or in known genomes to study structural variations. De novo sequence reads are preferably generated on those NGS platforms that generate longer sequence reads like Oxford Nanopore Technologies or PacBio SMRT; De novo sequencing can also be done on short read technologies like Illumina HiSeq platforms with different insert sizes (Paired end and mate pair).
Sequence reads from multiple platforms can be combined for Hybrid assembly (e.g., Illumina + PacBio Sequel) for cost effective coverage of large genomes. Sanger sequencing reads can also be used to complete the coverage.
AgriGenome offers Illumina HiSeq, MiSeq, Oxford Nanopore Technologies, PacBio Sequel and Sanger services for De novo genome sequencing and also Bioinformatics data analysis for efficient assembly, gene prediction and annotation.
For sales inquiries, please email us at: sales@aggenome.com
RNA-Seq is a powerful approach that enables researchers to discover, profile and quantify RNA transcripts in the whole transcriptome. As this method does not use predesigned probes or primers, it becomes an unbiased, hypothesis-free approach providing researchers with applications that are not possible by traditional microarray based methods.
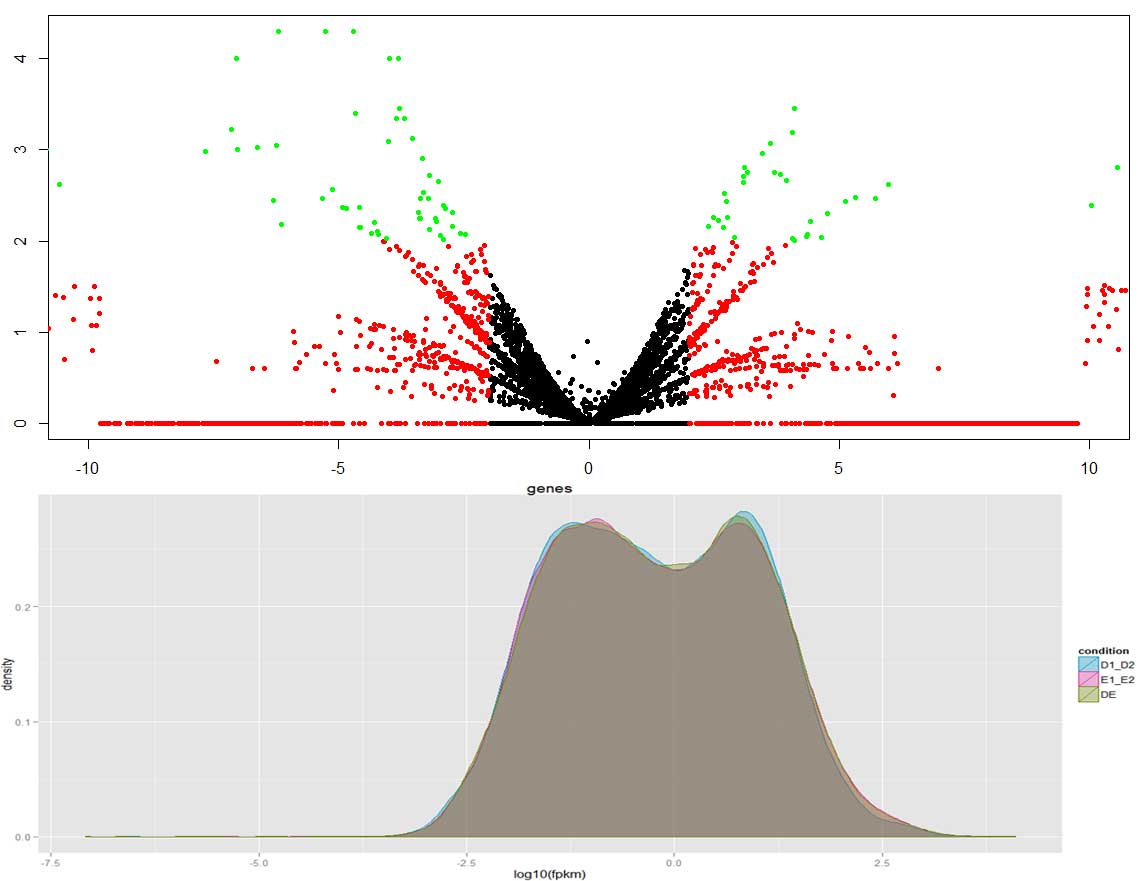
Basis for expression quantification in RNA-Seq
Platform : We offer Transcriptome sequencing on Illumina HiSeq platform
For sales inquiries, please email us at: sales@aggenome.com
Metagenomics, also known as Environmental genomics or Community genomics, is the study of multiple microbial species present in a specific sample that usually harbours complex microbial communities. Using NGS, it is possible to study microbial communities by sequencing the gene of choice (e.g. 16S, V3 and/or V4 regions) or the whole genome or transcriptome without the need for culturing.
Platform: Metagenomic sequence reads are generated on Illumina MiSeq/ HiSeq or PacBio Sequel depending on the experimental objectives. We can do detailed analysis of the metagenome NGS data.
For sales inquiries, please email us at: sales@aggenome.com
Small RNA sequencing is a powerful application enabling the discovery and profiling of microRNAs and other non-coding RNAs in any organism, without prior genome annotation. Using low RNA inputs, you can profile the differential expression of known microRNAs as well as detect novel microRNA targets and wide-ranging sequence variation or "iso-miRs" present in miRBase accessions.
With unprecedented sensitivity and dynamic range, small RNA sequencing methods allow for the most accurate detection and quantification of rare small RNA sequences.
Platform : We offer Transcriptome sequencing on Illumina HiSeq platform
For sales inquiries, please email us at: sales@aggenome.com
ChIP-seq is a technique or methodology to map transcription factor binding sites and histone modification status on a genome-wide scale. ChIP (Chromatin immunoprecipitation) technique comprises a few basic steps: cross-linking a protein to chromatin, shearing the chromatin, using a specific antibody to precipitate the protein of interest with its associated DNA, and reversing the cross linking and finally purifying the associated DNA fragments.
Once ChIP DNA (200-800bp in size) is submitted for sequencing, ChIP DNA libraries are made and run on Illumina MiSeq or HiSeq. This often generates tens to hundreds of millions of 50-100 bp sequences (also called short reads) from the 5' ends of ChIP-DNA fragments. We also do advanced data analysis of ChIP-seq data.
For sales inquiries, please email us at sales@aggenome.com
AgGenome Genotyping & Microarray Centre provides end-to-end services using the following platforms:
The cloning team provides customized cloning and sub-cloning services in a preferred choice of vector, starting from DNA, RNA or from the gene sequence.

The protein expression and purification services at AgriGenome provide a range of options to express and purify proteins of interest. Pure proteins are used in many applications in research and diagnostics.

AgriGenome offers production of high-affinity monoclonal reagent-grade Fabs and full-length IgGs for use in basic and clinical research labs. It uses licensed proprietary phage display libraries to generate synthetic Fabs/IgGs. The diversity of peptides displayed in the library is >1010, which guarantees high affinity Fabs against antigens from any source.

